1.7.13 Release Notes
Highlight of Features
Frontend
The Webclient configuration file has changed!
When upgrading to the v1.7.13 webclient, you will notice that i2b2_config_data.js has been renamed to i2b2_config_data.json. Your old configuration will still work with this new file name, but you will need to:
- remove all comments from the file (lines that begin with //).
- Escape slashes (e.g., / becomes \/)
There are also new optional parameters, documented below and in 1.4.2 Domain Configuration.
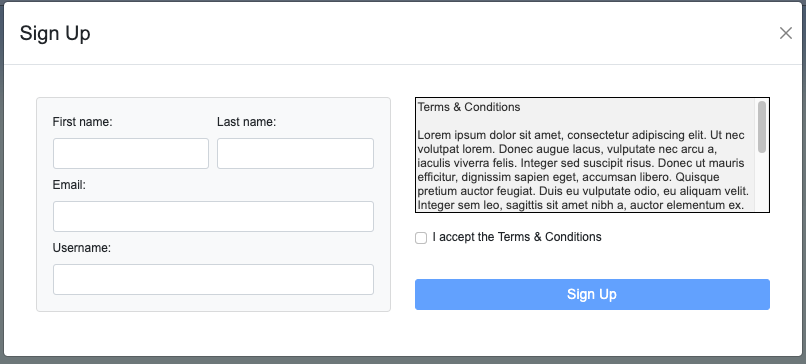
User Registration Tool
TODO
| registrationMethod | Y | String | NEW 1.7.13! Defines an information source for the new user registration tool. (If showRegistration is true, this parameter must be present.)
|
Community-Contributed Features
Contribution | Contributor | |
| SAML Authentication | Kevin Bui | i2b2 now includes support for SAML-based enterprise authentication via an institutional Identity Provider. See more information below. |
Ability to specify user parameter defaults | Michael Horvath | This change is meant to allowing user params to take precedence over hive params. Currently, it's the other way around. |
LDAP UPN Support | Michael Horvath Wake Forest | Active Directory enables other methods of binding which are more flexible besides just using the distinguished name. https://docs.microsoft.com/en-us/openspecs/windows_protocols/ms-adts/6a5891b8-928e-4b75-a4a5-0e3b77eaca52. This change is to enable binding the the User Principle Name form, which is very convenient when the distinguished names for users is not easily available (OU by department, etc.). |
| API to get all children of an ontology node | Kevin Bui | The metadata GetChildren API call, which returns information on the children of an ontology node, can now be configured to return multiple levels of children (e.g., children, children's children, etc.). This is done by specifying the numLevel parameters. By default, the function assumes numLevel = 1 and will return the direct descendants of the concept, which is one level of children. When the numLevel = -1 the function will return ALL descendants of the concept, otherwise the function will return up to and including the number of levels specified by numLevel (eg. numlevel=2 returns two levels of descendants, numLevel=4 returns four levels of descendants). |
| Totalnum Counter Performance Improvements | Darren Henderson University of Kentucky | Performance enhancements on SQL Server totalnum counting to not unnecessarily recompute temp tables. |
Backend Features
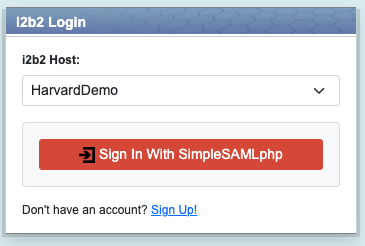
SAML Authentication
(LINKS DON'T WORK AND DOCUMENTATION IS INCOMPLETE.)
i2b2 now includes support for SAML-based enterprise authentication via an institutional Identity Provider. To configure this, you need to do the following:
Configure the webclient to use SAML:
loginType = "federated" see 1.4.2 Domain Configuration
configure SimpleSAMLPHP (now included with i2b2) to talk to your institution's Identity Provider. To set up SAML:
We will use SimpleSAMLphp for IdP. Place the following files to the folder /etc/shibboleth/:
If you would like to use your own IdP, please visit Configuration - Service Provider 3 - Shibboleth Wiki for advance configurations.
Place the following files in the directory /etc/httpd/conf.d/:
1) Setting up Apache and simplesamlphp: https://simplesamlphp.org/docs/latest/simplesamlphp-install.html
2) Configure the service provider and add an identity provider: https://simplesamlphp.org/docs/latest/simplesamlphp-sp.html
(You will need to generate a cert in /var/www/simplesamlphp/metadata/saml20-idp-remote.php)
Improved Totalnum Scripts
Totalnum Scripts have been updated to improve the Totalnum counter performance on both many ontology tables and very large(>1.5 million) ontologies such as ACT medications. Debug messages have also been added for troubleshooting and profiling purposes
Totalnum Scripts Setup
- In the Release_1-7/NewInstall/Metadata/ run the ant script to create the stored procedures.
ant -f data_build.xml create_metadata_procedures_release_1-7 - Run the stored procedures on your database. This can be done in two ways:
- Run the ant command to execute the data_build.xml file with below specified target
POSTGRESQL : ant -f data_build.xml db_metadata_run_total_count_postgresql
ORACLE : ant -f data_build.xml db_metadata_run_total_count_oracle
SQL SERVER : ant -f data_build.xml db_metadata_run_total_count_sqlserver - If using multiple fact tables, the recommended approach is to create a fact table view as the union of all your fact tables. (This is essentially going back to a single fact table, but it is only used for totalnum counting. This is needed to correctly count patients that mention multiple fact tables within a hierarchy.)
e.g., create view observation_fact_view as select * from CONDITION_VIEW union all select * from drug_view And then run the totalnum counter with the wildcard flag, to ignore multifact references in the ontology e.g., in SQL Server, exec RunTotalnum 'observation_fact_view','dbo','@','Y' Note this approach does not work if you have conflicting concept_cds across fact tables. - Execute the RunTotalNum stored procedure manually against your database in from a sql Client. This can take several hours. Example Usage:
- Run the ant command to execute the data_build.xml file with below specified target
Oracle:
begin
RUNTOTALNUM('observation_fact','i2b2demodata');
end;
-- You can optionally include a table named if you only want to count one ontology table (this IS case sensitive):
--begin
-- runtotalnum('observation_fact','i2b2demodata','I2B2');
-- end; Note: If you get the error as: ERROR at line 1: ORA-01031: insufficient privilege, then run the command:
grant create table to (DB USER)
SQL server:
exec RunTotalnum 'observation_fact','dbo','@' (the observation table name (for multi-fact-table setups), the schemaname, -- a single table name to run on a single ontology table, and a wildcard flag that will ignore multifact references in the ontology if 'Y')
PostgreSQL:
select RUNTOTALNUM('observation_fact','public')
– (replace 'public' by the schema name for the fact table)
– If using a schema other than public for metadata, you might need to run "set search_path to 'i2b2metadata','public' " first as well
- When finished, verify it is complete by checking that c_totalnum columns in your ontology tables contain numbers (not nulls).
These total counts will be visible in the ontology browser in the web client.
Additional New Stored Procedures
Age In Years Updater
When the CRC data is installed via ant, a new SQL script updates the age_in_years_num in the patient dimension based on the birth date.
As a reminder, this load process can be triggered with ant -f data_build.xml db_demodata_load_data in the CRC directory of NewInstall.
Concept Dimension Updater
Insert_Concept_FROMTableAccess is designed to populate concept_dimenison table using Table_access table records.
The stored procedure loops through the table_access and inserts values from corresponding c_table_name metadatatable which have
c_tablename column value as 'concept_dimension'
Example usage: exec Insert_Concept_FROMTableAccess
I2b2-Synthea data Load
A new option is now available for loading Synthea data files into i2b2. Synthetic patient data generated by Synthea is hosted on SyntheticMass..The Synthea sample files have been converted to i2b2-ACT format. The zipped data files can be downloaded from https://github.com/i2b2/i2b2-synthea
Synthea Load Process:
- All data sets (1k, COVID 10k, COVID 100k) have been verified to work EXCEPT the 100k patients in the large SyntheticMass Version 2 download. This version needs an extra step to delete invalid records before import. (Details coming soon.)
- Set up an i2b2 project with the ACT ontology.
- Run
create_synthea_table_<your dbServertype>.sqlin your project to create the Synthea tables. - Import the Synthea data you downloaded in step one into the Synthea tables in your project.
- Load the i2b2-to-SNOMED table in this repository into your project. https://www.nlm.nih.gov/healthit/snomedct/us_edition.html
- Click on the "Download SNOMED-CT to ICD-10-CM Mapping Resources" link to download. (You will need a UMLS account.)
- Unzip the file
- Import the TSV file into a table called SNOMED_to_ICD10 in your database.
- Run
synthea_to_i2b2_sqlserver.sqlto convert synthea data into i2b2 tables (this will truncate your existing fact and dimension tables!)- Replace references to
i2b2metadata.dboin the script. Use the database and schema where your ACT ontology tables are.
- Replace references to
ACT Version-4 Ontology data load
Metadata scripts are now available to load the latest ACT Version-4 Ontology into your i2b2 db schema
ACT4 data load process:
- Download the newinstall zip package from https://www.i2b2.org/software/download.html?d=452
- Extract the metadata\act folder from the downloaded zip folder
- Replace edu.harvard.i2b2.data\Release_1-7\NewInstall\Metadata\act folder with extracted new act folder
- Edit the db.properties file in your metadata folder to update the project properties to 'ACT'db.project=ACT
- From the edu.harvard.i2b2.data\Release_1-7\NewInstall\Metadata folder, run the ant command: ant -f data_build.xml db_metadata_load_data
- This will execute the SQL scripts form the edu.harvard.i2b2.data\Release_1-7\NewInstall\Metadata\act folder to create the new ACT4 Ontology tables
- and loads the corresponding metadata.
- You can now verify the new Ontology by logging into the webclient.
The following command will load the corresponding concept_dimension data of the Onbtology tables that will enable you to run queries in the webclient
From a sql Client>select 'insert into concept_dimension select c_dimcode AS concept_path, c_basecode AS concept_cd, c_name AS name_char, null AS concept_blob, update_date AS update_date, download_date as download_date, import_date as import_date, sourcesystem_cd as sourcesystem_cd, 1 as upload_id from '
+c_table_name+' where m_applied_path=''@'' and c_tablename=''CONCEPT_DIMENSION'' and c_columnname=''concept_path'' and c_visualattributes not like ''%I%'' and (c_columndatatype=''T'' or c_columndatatype=''N'') and c_synonym_cd = ''N'' and (m_exclusion_cd is null or m_exclusion_cd='''') and c_basecode is not null and c_basecode!='''''
from <your_dbschema>.dbo.TABLE_ACCESS where c_visualattributes like '%A%'
Security Enhancements
- i2b2 has been made more secure by addressing parameterization and other potential vulnerabilities according to a Veracode scan.
- Log4J has been upgraded to the latest version.
Improved db Upgrade Process
Currently i2b2 db upgrade is a multi-step process of running upgrade scripts and stored procedures. This release provides a set of upgrade scripts which will perform the complete db upgrade.
based on your current build version.
For example: Following Ant command will upgrade your db instance from 1.7.09c to latest version.
>ant -f data_build.xml upgrade_table_release_1-7-09c upgrade_table_release_1-7-10 upgrade_table_release_1-7-11 upgrade_table_release_1-7-12
Steps to Perform db upgrade:
- Backup your existing data folder
- Copy all the folders from the extracted download data folder into your existing data Upgrade folder
Example: Downloads\2b2core-upgrade-1712a\i2b2\data to C:\opt\edu.harvard.i2b2.data\Release_1-7\Upgrade\. This will replace
existing Crcdata, Hivedata, Metadata, PMdata folders.
Alternative to above step, navigate to the edu.harvard.i2b2.data\Release_1-7\Upgrade\ directory of your extracted folder - Copy the db.properties files from your back up into the respective locations(namely Crcdata, Hivedata, Metadata, PMdata )
- Open the command prompt and navigate to cell folders and run the following upgrade ant commands on your i2b2 database instance, where {db} can be Oracle, sqlserver or postgresql.
Alternative to above Step, you can run individual SQL scripts on your db instance in place of ant commands.
In data folder\Release_1-7\Upgrade\ run the ant commands under each individual cell subfolder as below. | |
| Upgrade From Build | Upgrade to Latest build |
|---|---|
1.7.09c | In the Crcdata folder run the following ant command: ant -f data_build.xml upgrade_table_release_1-7-09c upgrade_table_release_1-7-10 upgrade_table_release_1-7-11 upgrade_table_release_1-7-12 In the Hivedata folder run the following ant command: ant -f data_build.xml upgrade_hive_tables_release_1-7-09c upgrade_hive_tables_release_1-7-10 upgrade_hive_tables_release_1-7-11 upgrade_hive_tables_release_1-7-12 In the Metadata folder run the following ant command: ant -f data_build.xml upgrade_tables_release_1-7-09c upgrade_tables_release_1-7-10 upgrade_tables_release_1-7-11 upgrade_tables_release_1-7-12 In the PMdata folder run the following ant command: ant -f data_build.xml upgrade_pm_tables_release_1-7-09c upgrade_pm_tables_release_1-7-10 upgrade_pm_tables_release_1-7-11 upgrade_pm_tables_release_1-7-12 |
1.7.10 | In the Crcdata folder run the following ant command: ant -f data_build.xml upgrade_table_release_1-7-10 upgrade_table_release_1-7-11 upgrade_table_release_1-7-12 In the Hivedata folder run the following ant command: ant -f data_build.xml upgrade_hive_tables_release_1-7-10 upgrade_hive_tables_release_1-7-11 upgrade_hive_tables_release_1-7-12 In the Metadata folder run the following ant command: ant -f data_build.xml upgrade_tables_release_1-7-10 upgrade_tables_release_1-7-11 upgrade_tables_release_1-7-12 In the PMdata folder run the following ant command: ant -f data_build.xml upgrade_pm_tables_release_1-7-10 upgrade_pm_tables_release_1-7-11 upgrade_pm_tables_release_1-7-12 |
1.7.11 | In the Crcdata folder run the following ant command: ant -f data_build.xml upgrade_table_release_1-7-11 upgrade_table_release_1-7-12 In the Hivedata folder run the following ant command: ant -f data_build.xml upgrade_hive_tables_release_1-7-11 upgrade_hive_tables_release_1-7-12 In the Metadata folder run the following ant command: ant -f data_build.xml upgrade_tables_release_1-7-11 upgrade_tables_release_1-7-12 In the PMdata folder run the following ant command: ant -f data_build.xml upgrade_pm_tables_release_1-7-11 upgrade_pm_tables_release_1-7-12 |
| 1.7.12 | In the Crcdata folder run the following ant command: ant -f data_build.xml upgrade_table_release_1-7-12 In the Hivedata folder run the following ant command: ant -f data_build.xml upgrade_hive_tables_release_1-7-12 In the Metadata folder run the following ant command: ant -f data_build.xml upgrade_tables_release_1-7-12 In the PMdata folder run the following ant command: ant -f data_build.xml upgrade_pm_tables_release_1-7-12 |
Changelog
Database Drivers
The JDBC drivers were updated to the following versions.
Driver | ojdbc8.jar | postgresql-42.2.5.jar | mssql-jdbc-7.0.0.jre8.jar |
|---|---|---|---|
New Version | Oracle 12.2.0.1 | PostgreSQL 42.2.5 | MS Sql Server 7.0.0 |
Supported Db Server versions
Server Type | SQL Server | Oracle | Postgres |
|---|---|---|---|
Supported Version/s | 2012+ (tested with up to 2019) | 12g+ and 21c | 9 to 14 |
Supported software versions
Application Type | Java | Wildfly | Apache HTD | Apache Ant | Apache Axis2 | PHP |
|---|---|---|---|---|---|---|
Supported Version/s | 8 or 11 | 17.0.0 | 2.4.17 or higher | 1.9.6 | 1.7.1 | 7.2.27 or higher |
i2b2 Database Changes
| New db updates |
|---|
|
i2b2 Server and Client Changes
New Features and Improvements
| Webclient | Core-server |
|---|---|
|
|
Bug Fixes
| Webclient | Core-server |
|---|---|
|
|
Notes for Developers
For Java 11 install, if you change xsd, then modify the gensource.