- Dashboard
- Health Ontology Mapper
- HOM Home
- Attachments
- Translational_Bioinformatics_Warehouse_Database_Design_v4.doc
Translational_Bioinformatics_Warehouse_Database_Design_v4.doc
The CTSA Ontology Mapper and Discovery Suite:
A Rules-Based Approach to Integrated Data Repository Deployment
Translational Data Warehousing Design
Strategies for Supporting the Ontology Mapper Project
1University of Pennsylvania – Health System
2University of California , San Francisco – Academic Research Systems
3University of Rochester - Medical Center
4University of California , Davis - Clinical & Translational Science Center
Contents
3.1 Structured Database Design
3.2 Dimensional Database Design
3.3 Integrated Data Repository Design
3.4 Federated vs. Centralized Approach
3.4.1 Bridging data across systems made easier by centralization
4. Example Data Warehousing Design Approaches
5. Mapper Database Design - Extending i2b2
Figures
Figure 1 - Data Warehouse Architecture - (Inmon et al., 2001)
Figure 2 - Data Warehousing Components (Kimball, 2002)
Figure 3 - Normalized Data Model Example
Figure 4 - Star Schema Example (Kimball, 2002)
Figure 5 - Complex data governance (top) can be exchanged for rules encoding (bottom)
Figure 7 - i2b2 Database Design
Figure 8 - Ontology Mapper Schema Design
Revision History
|
Date |
Version |
Reason For Changes |
Modified By: |
|
5/30/08 |
0.1 |
Initial draft |
Marco Casale |
|
|
|
|
|
|
7/16/08 |
0.2 |
Added Prakash Lakshminarayanan’s work on Mapper Database Design. Updated Core i2b2 schema design |
Marco Casale
|
|
8/21/08 |
0.3 |
Updated Prakash’s latest mapper database design approach. Inserted IDR database design approach from Rob Wynden |
Marco Casale |
|
|
|
Inserted Clinical value of IDR approach from Dr. Mark Weiner Inserted i2b2 core star schema design and description |
|
The Ontology Mapper Project is intended to provide a mechanism for enhancing translational bioinformatics. It is a general purpose instance mapper which can be used to translate local data, which may not be encoded according to any standard format, into one or more formal encodings. This project takes a different approach to the standard data warehousing model and enhances it with the use of ontology mapping for biomedical terminology. In order to accommodate this novel approach, a review of current data warehousing design will be compared and contrasted.
|
Term |
Definition |
|
IDR |
Integrated Data Repository |
|
ETL |
Extract, Transform and Load |
|
HL7 |
Health Level 7 – Standard used for information transportation amongst disparate IT systems |
|
i2b2 |
Informatics for Integrating Biology and the Bedside |
|
ODS |
Operational Data Store |
|
CIF |
Corporate Information Factory |
|
Fact |
A business measure |
|
Dimension |
Textual descriptions of the business. Descriptions of facts |
Data warehousing is a method used to convert data into information. A clinical data warehouse can provide a structure for organizing data and render a positive impact for the purpose of patient care, biomedical research and biomedical education. However, developing a data warehouse of clinical, clinical research and bench science data (called an Integrated Data Repository or IDR) encompasses many difficult challenges. Such a data warehouse cannot be designed in one simple step.
The data warehouse architecture model shown in Figure 1 depicts the process of transforming operational data into information for the purpose of generating knowledge within an organization. The diagram displays data flowing from left to right in accordance with the corporate information factory (CIF) approach (Inmon et al, 2001). According to Inmon, the data enters the CIF as raw data collected by operational applications. The data is transformed through extract, transform and load processes and stored in either the data warehouse or the ODS, operational data store. “Often up to 80 percent of the work in building a data warehouse is devoted to the extraction, transformation, and load (ETL) process: locating the data; writing programs to extract, filter, and cleanse the data; transforming it into a common encoding scheme; and loading it into the data warehouse.” ( Hobbs , Hillson & Lawande, 2003, 6)
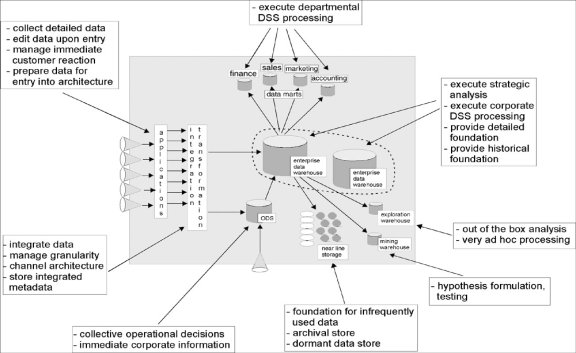
Figure 1 - Data Warehouse Architecture - (Inmon et al., 2001)
According to Kimball (2002), the data warehousing architecture can further been delineated into four categories:
- Source Systems
- Data Staging Area
- Data Presentation Area
- Data Access Tools
Source Systems may be composed of operational systems, legacy based systems or other data repositories. Data is transmitted or extracted from the source systems into the data staging area where it is processed. It should be noted that the staging area is not intended for direct query access. The purpose of data processing is to normalize the data, using standard data types, data lengths, precision of data and data naming conventions amongst the inbound extractions.
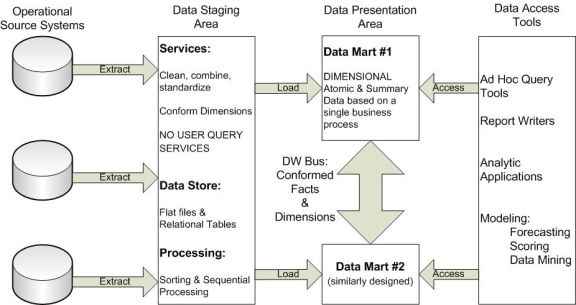
Figure 2 - Data Warehousing Components (Kimball, 2002)
Furthermore, after the data has been prepared, it is loaded into the de-normalized schema of the data warehouse or data marts and resides there in a fine grain level of detail. The logical design of a data warehouse is usually composed of the star schema. “A star schema is a simple database design (particularly suited to ad-hoc queries) in which dimensional data (describing how data are commonly aggregated) are separated from fact or event data ( describing individual business transactions).” (Hoffer, Prescott & McFadden, 2002, 421) The data mart design is intended to support the data presentation layer.
The final stage is in the process of converting data into information through reporting, analysis and data mining techniques. (Inmon et al, 2001) Kimball describes this stage as the data access tools area or in modern day vernacular, the business intelligence layer.
There are a variety of data modeling approaches for data warehousing design. These approaches can range from highly structured database design, dimensional database design, entity, attribute, value design, object oriented design to name a few.
3.1 Structured Database Design
Operational transaction processing systems are commonly created using a highly structured or normalized data model. This model is intended to reduce several anomalies (insert, update and delete) that may occur during data transaction processing. The goal of normalization theory, developed by E.F. Codd and CJ Date, is to determine where the functional dependencies exist within and amongst relations. It is supported by mathematical set theory, relational algebra and relational calculus. The end result is to produce a data model which reduces the opportunity for data redundancy, and provides the highest level of data consistency. Figure 3 is a depiction of a third normal form data model of patient visits and their associated observations.
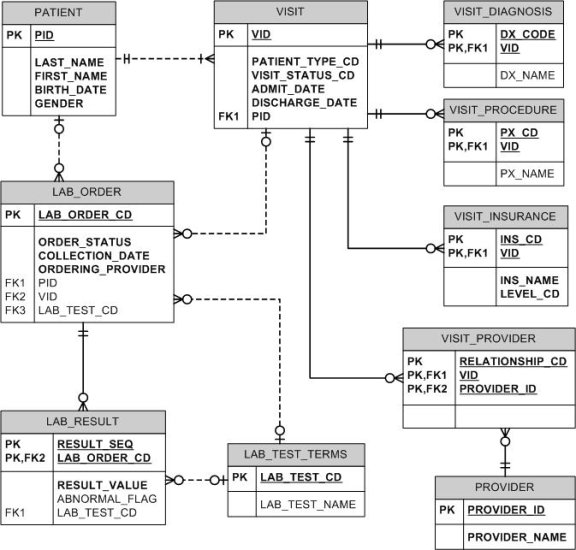
Figure 3 - Normalized Data Model Example
3.2 Dimensional Database Design
In contrast, Kimball proposed a dimensional modeling approach for data modeling design in support of high performance queries rather than high volume transaction processing. The dimensional model is composed of a fact table joined to several dimensions. A fact table is the primary table of the dimensional model. Each fact table contains measurable facts. “A row in a fact table corresponds to a measurement. A measurement is a row in a fact table. All the measurements in a fact table must be at the same grain.” (Kimball, 2002). Figure 4 is an example of a dimensional model that illustrates a fact table of ‘Daily Sales’ related to dimensions of Date, Product and Store.

Figure 4 - Star Schema Example (Kimball, 2002)
“The dimension tables contain the textual descriptors of the business.” (Kimball, 2002). Each dimension table may be composed of many attributes. The dimension attributes are used to help qualify the criteria used in generating queries from the star schema. Every dimension table contains a primary key. Furthermore, the fact table contains a foreign key to every dimension table and typically, the primary key of the fact table is a composition of all of the foreign keys in the fact table, a composite primary key.
Now that the transformed data resides in the underlying fact and dimension tables of the data warehouse, the next step entails deriving the necessary aggregates that will define the associated data marts. “An aggregate is simply the rollup of an existing fact table along one of its dimensions, which more often than not is time.” (Scalzo, 2003, 147) For example, if a fact table holds daily automobile fatalities, then location-based aggregates might be town, county, and state. Thus, since the overall purpose of a data warehouse is to support the online analytical processing (OLAP) capabilities associated with data mining, the data in the warehouse must be propagated, as referred to in Figure 1, into information residing within data marts.
Data marts serve to hold specialized, codified, dimensional, aggregated data roll-ups in support of the OLAP tools that will be used to discover business relationships and organization insights. However, slicing a cube does not always yield the desired knowledge. As a result, there are times when exploratory data mining processes must be initiated against the data warehouse itself, as opposed to a specialized multidimensional array. This drill down technique can also be accomplished with OLAP tools, specialized data mining software or even customized ad-hoc programmatic queries running against the finer level of detail. The data access tools in Figure 2, processes all of these requests and coordinates the communication exchange between the warehouse or marts and the end user’s tools.
3.3 Integrated Data Repository Design
A complex problem exists within data warehousing design. It is typically found in the extract, transformation and load processes that require data governance. The data governance’s role is to determine how the data will be interpreted and managed within the data warehouse. This phase of the data warehouse is encumbered with inflexibility and an exhaustive use of resources.
The integrated data repository (IDR) approach shifts this responsibility by incorporating an ontology mapping software service which runs inside of the IDR. This service provides the capability to map data encoded with different ontologies into a format appropriate for a single area of specialty, without preempting the further mapping of that same data for other purposes.
This approach would represent a fundamental shift in both the representation of that data within the IDR and a shift in the manner in which resources are allocated for servicing Translational Biomedical Informatics environments. Instead of relying on inflexible, pre-specified data governance, the proposed model instead shifts resources to the handling of user requests for data access. Data interpretation happens as a result of a specific researcher request, and therefore only as it is deemed useful.
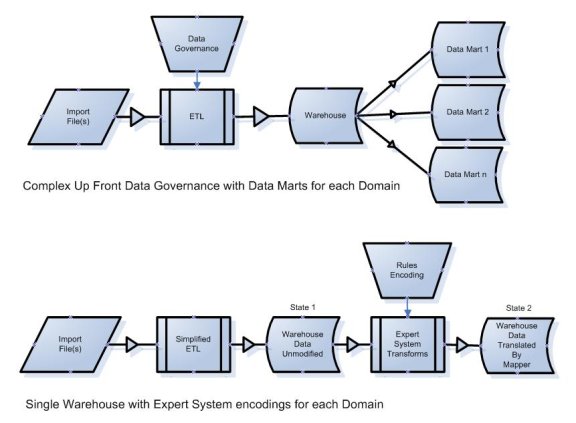
Figure 5 - Complex data governance (top) can be exchanged for rules encoding (bottom)
The data in the bottom diagram will exist in both State 1 and State 2 simultaneously. The source data is always maintained within the warehouse in its source format as well as in its translated formats. Data may exist only once in State 1, but that same data may be translated multiple times and therefore may exist in several different formats in State 2. This is consistent with Theme 2 above which seeks to enable the organization of the same data using differing hierarchies and terminology encodings.
3.4 Federated vs. Centralized Approach
The debate regarding a federated data warehouse design versus an ‘all in one’ centralized data warehouse design has been a key point between the two main data warehousing factions between Bill Inmon and Ralph Kimbal. The following example outlines the differences for the two approaches and justifies why the centralized approach is the favorable choice in biomedical informatics.
3.4.1 Bridging data across systems made easier by centralization
Many institutions have electronic clinical data that is decentralized across different departments. That infrastructure can be used to create an integrated data set with information that spans the source systems. However, the decentralization creates the need for redundant steps, lengthy iterative processes to refine the information, and requires that more people have access to protected health information in order to satisfy the research information needs. To illustrate these issues, the following describes the workflow needed to define a cohort of people known to have cardiovascular disease and laboratory evidence of significant renal disease defined by an elevated serum creatinine.
In the decentralized system, where should the investigator start? He can begin by going to the billing system that stores diagnoses and get a list of PHI (Personal Health Information) of people with a history of a heart attack. Then he can transport that list of identifiers to the people who work in the laboratory and request the serum creatinine levels on that set of patients, and then limit the list to those who have an elevation. The lab will have to validate the patient matches generated by the billing system by comparing PHI, a step redundant with the billing system. Furthermore, many of the subjects associated with heart attack may not have the elevated creatinine, so, in retrospect, the PHI of these people should not have been available to the people running the query in the lab. Perhaps the cohort that was generated was not as large as expected, and the investigator decides to expand the cohort to those patients with a diagnosis of peripheral vascular disease and stroke. He then has to iterate back to the billing system to draw additional people with these additional diagnoses, and then bring the new list of patient identifiers to the lab to explore their creatinine levels.
The centralized warehouse as proposed will conduct the matching of patient identifiers behind the scenes. The information system will conduct the matching of patients across the different components of the database, so that identifiers do not have to be manually transported and manipulated by the distinct database managers at each location. The result is that a centralized warehouse is radically more secure than a decentralized warehouse due to the reduced exposure of PHI. Further, if the query produces results that are not satisfactory, the cycle of re-querying the data with new criteria will be faster, and user controlled.
4.1 Partners Healthcare RPDR
A significant challenge for applications of clinical data warehousing has been the ability to make them facile for non-database experts. Typically, database experts are required to write complex query statements in order to extract data. The following example depicts how one organization dealt with this difficult dilemma.
Partners Healthcare Inc. created a research patient data repository (RPDR) in support of clinical research. Partners recognized the need for several database experts writing complex queries, when they started to receive 100-200 query requests per week. Partners decided to develop a graphical analytic tool to help novice database users create effective mining algorithms. (Murphy et al, 2003) Partners set out to develop a Querytool for the RPDR shown in Figure 5. However, they were confronted with many challenges.
Murphy described the first challenge as allowing novice population of computer users to navigate the metadata and formulate the queries without computer expert intervention. The Querytool leverages medical vocabularies such as ICD9, SNOMED-CT, LOINC, NDC and CPT4. Unfortunately, each vocabulary presents its own obstacles. For instance, ICD9 codes contain categories that can be used respectively for coding. On the other hand, NDC drug codes can be reused so that their meanings change from year to year. (Murphy et al).
The second challenge involved construction logic. “Patient inclusion and exclusion criteria for most research studies are jumbles of different types of information.” (Murphy et al). For instance, a researcher might request all patients who have been diagnosed with a certain disease and are in a particular age group but do not take a certain medication. Therefore, it was the Querytool’s objective to disguise the complex construction logic with an intuitive interface (Figure 5).
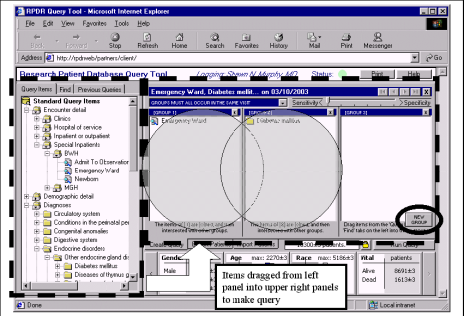
In short, the Querytool (now referred to as The Workbench) is an excellent example of a data warehouse graphical user interface to novice database users for clinical research. The end user is easily able to narrow down the 1.8 million patients at Partners to a manageable number between 100-1000 patients. Finally, Murphy states “The Querytool has achieved excellent acceptance at Partners Healthcare Inc. and is the most prevalent way of obtaining research cohorts.”
The Partners Healthare RPDR system has since be released into open source as the i2b2 (Informatics for Integrating Biology and the Bedside) integrated data repository platform. The i2b2 platform is the first open source general purpose integrated data repository system intended for use within translational medical informatics.
4.2 Partners Healthcare i2b2
The following figure depicts the core i2b2 database schema. It conforms to a star schema design based on Ralph Kimball's initial work. Figure 8 is a depiction of the core i2b2 data model design. Its purpose is to support a clinical research cell (CRC) in the i2b2 framework, which holds data such as clinical trials, medical records systems and many other heterogeneous biomedical data sources.
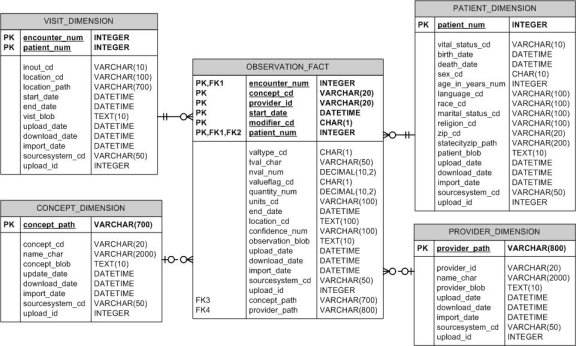
Figure 7 - i2b2 Database Design
The star schema is composed of a fact table, the observation_fact, and four dimension
tables: visit_dimension, patient_dimension, provider_dimension and concept_dimension.
In healthcare, a logical fact is an observation on a patient. It is important to note that an observation may not represent the onset or date of the condition or event being described, but instead is simply a recording or a notation of something. For example, the observation of ‘diabetes’ recorded in the database as a ‘fact’ at a particular time does not mean that the condition of diabetes began exactly at that time, only that a diagnosis was recorded at that time (there may be many diagnoses of diabetes for this patient over time).. (Murphy, 2008)
The fact table contains the basic attributes about the observation, such as the patient and provider numbers, a concept code for the concept observed, a start and end date, and other parameters described below in this document. (Murphy, 2008) Furthermore, prior to the introduction of The Ontology Mapper, the observation_fact table was composed of a composite primary key consisting of six attributes. The combination of these six attributes uniquely identifies a record in the fact table.
The dimension tables support the fact table by providing an opportunity to view facts from different perspectives using selection criteria. The Patient_Dimension contains a record for each patient in the database. The patient attributes are aggregated based on an algorithm that examines the most frequent value over time. The provider dimension supports a detailed description of clinical care providers who have a relationship with the patient and the patient’s observations. The visit_dimension is a list of all of the visits or encounters for the set of patients in the database. Finally, the concept_dimension contains a record for the unique clinical terms and their hierarchical structure based on the concept_path field.
The design of the i2b2 schema represents a further evolution of traditional fact table design. Each row in the fact table is not simply described with a concept code but also with a concept path. These concept paths can be traversals of formal ontologies or they can be ad-hoc paths which reflect of the structure of source software systems. As new source systems are imported and as ontologies are updated, new paths are simply added to the concept dimension table. Both older deprecated and newly added paths are permanently retained within the concept dimension table.
This is an architecture first proposed by Jim Cimino.
[r1]
4.3 Ontology Mapper Database Design - Extending i2b2
To support the alternate data governance model shown in Figure 5 above a further evolution of the database model provided by i2b2 was required. In order to support this refinement of the data management process the ability to map data (instance mapping) after ETL must be added to the data warehousing model. The proposed Ontology Mapper functionality that enables the creation of maps for the transformation of data from source encoding to target encoding required several changes on the original i2b2 schema. These changes that were effected on the i2b2 schema are outlined herewith on a table by table basis.
In order to maintain data integrity, a design approach was made to diminish the alteration of the core i2b2 star schema design. Minimal changes were made to the observation_fact table to lesson any impact on the primary key. These changes support the relationship between the core fact and the instance map that was used in support of that fact. In general, the primary key is necessary for providing a unique instance of an observation. In addition, all fields in a record should have significance. For instance, if an aggregation of records are generated across a particular value, the primary key must continue to uniquely identify the instance of the aggregated record. Another approach would be to create an aggregated_observation_fact table based [r2] on applied aggregations.
Furthermore, the reduction of alterations to the star schema will help to reduce changes incurred by subsequent releases of the i2b2 software. As each database upgrade is applied, the following minor modifications will also need to be applied.
ENCODING_DIMENSION
This is a new table created for the purpose of storing the various encodings used in the Mapper system. ENCODING_CD serves as the primary key for the table. Other columns of the table are enlisted below:
- ENCODING_NAME – the name of the encoding
- ENCODING_DESC – description of the encoding
- CREATE_USER – the user that created the encoding
- CREATE_DATE – the date of creation of the encoding
- UPDATE_USER – the user that last updated the encoding
- UPDATE_DATE – the date of last update of the encoding
MAP_DIMENSION
This table has been created to store info on the instance mapper xml files uploaded onto the system. MAP_ID which is a running sequence no. serves as the primary key for this table. Other columns of this table are enlisted below:
- MAP_NAME – the name assigned for this map
- CONCEPT_PATH – the concept path for which this map applies
- SOURCE_ENCODING – the source encoding of the map
- TARGET_ENCODING – the target encoding of the map
- MAP_PATH – the physical path of the mapper xml instance in the system
- MAP_DESC – description of the map
- IMPORT_DATE – the date of import of the map into the system
- UPDATE_DATE – the date of last update of the map
- UPLOAD_USER – the user that uploaded the map onto the system
- UPDATE_USER – the user that last updated the map
- LAST_RUN_DATE – the last execution date of the map
OBSERVATION_FACT
- Added column CONCEPT_PATH referenced from table CONCEPT_DIMENSION for storing the concept path pertaining to the concept code used in this table. This is required since the original i2b2 design supports only the CONCEPT_CD (this can be the same for multiple concept paths) and resolving the concept path from the same is difficult.
- Added column ENCODING_CD referenced from table ENCODING_DIMENSION to denote the encoding type this record is encoded with.
- Column OBSERVATION_FACT_ID added to store a running sequence no. for uniquely identifying each record in the table.
MAP_DATA_FACT
- This is a new table designed to store the records created as a result of Ontology Mapper execution. All the transformed records in target encodings will be housed in this table.
- This table has been created to segregate the transformed data from the source data and also to keep the i2b2 fact table (OBSERVATION_FACT) design intact. The Ontology Mapper functionality required several design changes on the OBSERVATION_FACT which might have rendered this table incompatible with future i2b2 releases. This new table resolves the aforesaid issue by providing the flexibility of effecting new design changes required for the Mapper functionality.
- In accordance with Mapper functionality the new table is designed to have a new primary key combination consisting of columns CONCEPT_PATH, ENCODING_CD, MAP_ID and OBSERVATION_FACT_ID.
- CONCEPT_PATH column is referenced from CONCEPT_DIMENSION to indicate the concept path for which this map applies.
Please note: we are therefore further extending an architecture first proposed by Jim Cimino which utilizes a concept dimension table where all paths are stored regardless of whether they represent currently accepted paths within a formal ontology. All paths are simply retained as new paths are created and old paths are deprecated. In this extension to that algorithm, we are also representing newly instance mapped data as yet more paths within that same concept path table.
- MAP_ID column of this table is referenced from MAP_DIMENSION and stores info on the map that created this record.
- ENCODING_CD is referenced from ENCODING_DIMENSION and holds info on the encoding used in creation of this record.
- OBSERVATION_FACT_ID column is used to refer back source record of OBSERVATION_FACT from which this record has been generated.
- MAP_DATE column is designed to store the record creation date.
- All other columns in this table resemble those in the OBSERVATION_FACT both in name and purpose. Only few columns from the OBSERVATION_FACT have been retained so as to support aggregates and archetypes [r3] . In other words only columns on which aggregation can be performed have been retained.
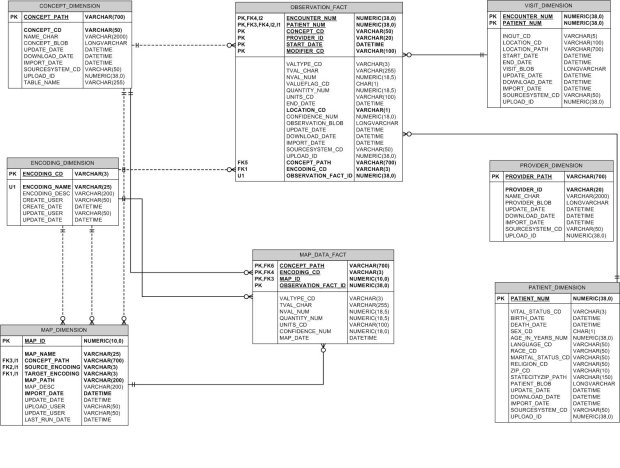
Figure 8 - Ontology Mapper Schema Design
These alterations to the i2b2 schema produced a system which did interoperate with the i2b2 schema but which would have required many alterations to the existing i2b2 application server source code. Therefore the following set of modifications was also made which provide a tighter coupling of the Ontology Mapper to the i2b2 application server.
4.3 Ontology Mapper Database Design - i2b2 Data Mart Generation
This final set of modifications made to i2b2 was required in order to provide compatibility with the existing i2b2 application server code. Additionally these modifications also offer a fundamental improvement in i2b2 security, storage requirements and performance.
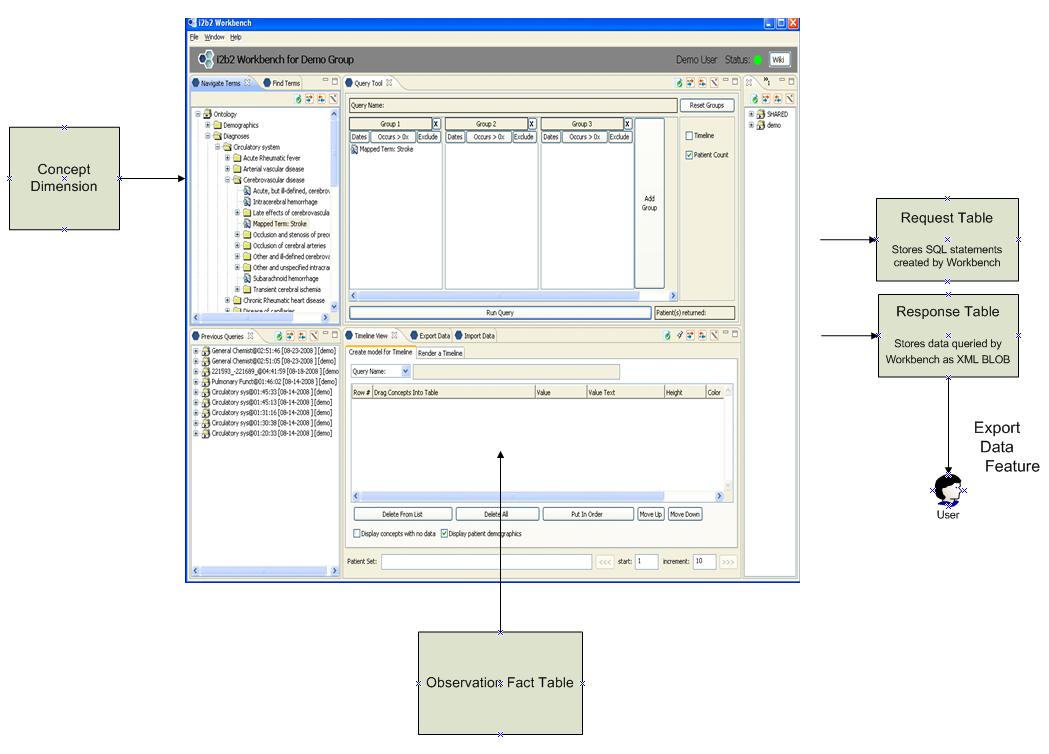
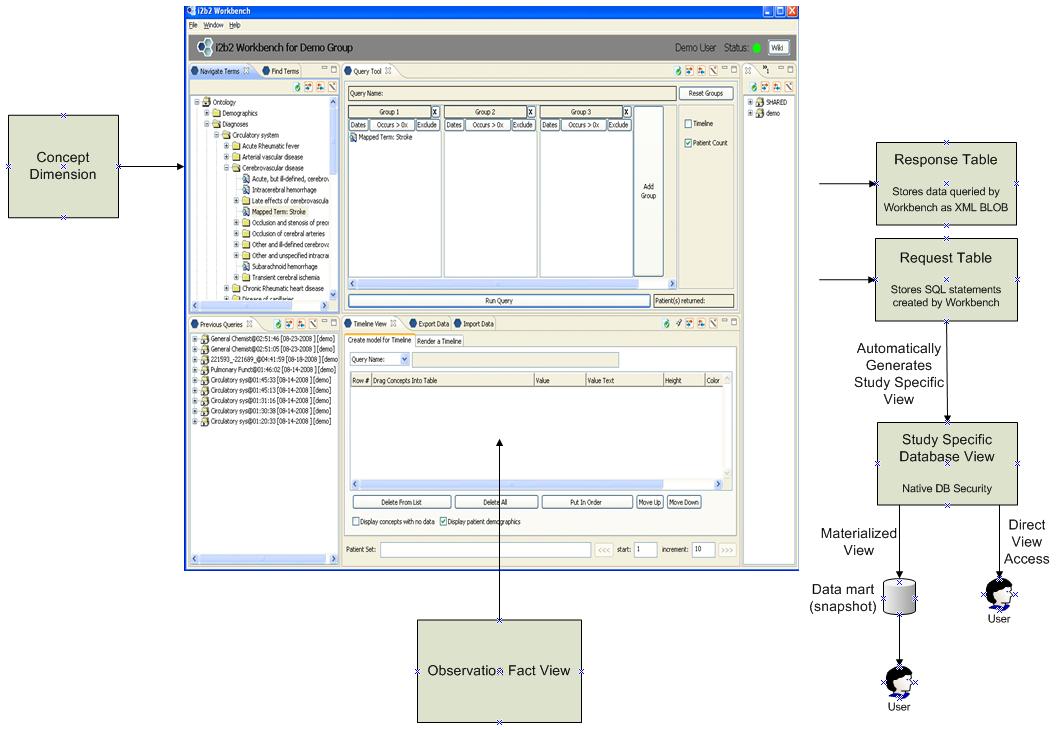
Figure 9 – i2b2 Workbench and related tables
In the current i2b2 workbench software the concept dimension is used to populate the list of concept paths displayed on the left of the screen. The user generates a query via a drag-and-drop of these concept paths into the query boxes at the top right. The query results are populated from the observation fact table using the box on the lower right.
The SQL query which is generated by this UI is stored in the request table. The actual data returned is stored within the response table as an XML blob. Using an i2b2 feature called the Export Data Feature the user is then granted access to the data contained within that response data BLOB.
This architecture has several disadvantages:
1) Data stored within the observation fact table is replicated into the response data BLOB. For users which require a snapshot of data this format may be acceptable. However database BLOB fields have size limitations and we expect researcher queries to require access to very large sets of IDR data. Replication of such large datasets has distinct disadvantages regarding the efficient usage of storage space.
Also not all users want to have access to information as snapshots. The Ontology Mapper will continuously instance map incoming data which is presented to the system during regular ETL processing. Some users will request access to this newly instance mapped data as soon as it becomes available. An algorithm which is based on
2) This architecture requires to maintenance of a duplicate security paradigm outside of the host database environment. The BLOBs which are created and exported to the researcher must be permissioned for usage solely by the researcher’s study staff under IRB (Institutional Review Board) protocol. Since this mechanism is not using native database security for data delivery that security mechanism must be replicated within the application server layer.
Security models which are dependent on application server security are inherently less secure than models which leverage native database security. In models which leverage database security, if the application server layer is hacked then the would-be hacker would still not obtain access to the data in question.
3) MARK – ADD THIRD CASE HERE [r4]
An explanation of the Mapped Term prefix and justification for such may go well here………
To address these concerns and to provide an easier method of integration of the Ontology Mapper into the i2b2 framework we have therefore made the following additional modifications to the i2b2 schema.
Figure 10 – Extensions to i2b2 to support the generation of Study Specific Views
Please note that in this model instead of providing access to end users via the XML BLOB of response data (referred to as the i2b2 CRC or Clinical Research Chart), the user is instead granted access automatically to a database view. That database view can then be either manipulated directly or it can be materialized into a data mart when necessary.
Also the original workbench software would be modified so that the data shown in the lower right portion of the screen is derived from an Observation Fact View which is comprised of data from both the Observation Fact Table and the Mapped Data Fact Table.
The Observation Fact View is created such:
CREATE
SELECT OBS.* FROM
OBS_FACT, MAPPED_DATA_FACT
WHERE OBS.FACT.OBSERVATION_FACT_ID = MAPPED_DATA_FACT.OBSERVATION_FACT_ID
Conclusion
References
Adelman, Sid & Terpeluk-Moss, Larissa (2000). Data Warehouse Project Management . Boston : Addison Wesley.
Berry , M. J. A. & Linoff, G. (2000). Mastering Data Mining: The Art and Science of Customer Relationship Management . Hoboken : John Wiley & Sons.
Berson, A. & Smith, S. & Thearling, K. (2000). Building Data Mining Applications for CRM. New York : Computing McGraw-Hill.
Bray, T. (2001). What is RDF
Referenced July 26, 2003 from http://www.xml.com/pub/a/2001/01/24/rdf.html?page=2
Delmater, R. & Hancock, M. (2001). Data Mining Explained: A Manager's Guide to Customer-Centric Business Intelligence. St. Louis : Digital Press.
English, Larry P. (1999). Improving Data Warehouse and Business Information Quality. New York : Johm Wiley & Sons.
Han, Jiawei & Kamber, Micheline (2001). Data Mining: Concepts and Techniques. USA : San Francisco : Morgan Kaufmann.
Hobbs , Lilian; Hillson, Susan & Lawande, Shilpa (2003). Oracle9iR2 Data Warehousing. Burlington : Digital Press.
Hoffer, Jeffrey A.; Prescott, Mary B. & McFadden, Fred R. (2002). Modern Database Management. Upper Saddle River : Prentice Hall.
Inmon, W. H. (2001)(a). A Brief History of Metadata
Retrieved July 20, 2003 from http://www.inmoncif.com//library/whiteprs/wp.asp#Metadata
Inmon, W. H. (2001)(b). An Illustrated Taxonomy of Metadata
Retrieved July 20, 2003 from http://www.inmoncif.com//library/whiteprs/wp.asp#Metadata
Inmon, W. H. (2002). Building the Data Warehouse.
New York : John Wiley & Sons, Inc.
Inmon, W. H., Imhoff, C., Sousa, R. (2001). Introducing the Corporate Information
Factory 2nd Edition New York : John Wiley & Sons, Inc.
Inmon, W. H. (2000). Metadata in the Data Warehouse
Retrieved July 20, 2003 from http://www.inmoncif.com//library/whiteprs/wp.asp#Metadata
Inmon, W. H. (1997). Metadata in the Data Warehouse: A Statement of Vision
Retrieved July 20, 2003 from http://www.inmoncif.com//library/whiteprs/wp.asp#Metadata
Jarke, Matthias; Maurizio, Lenzerini; Vassiliou, Yannis & Vassiliadis, Panos (2003). Fundamentals of Data Warehouses. (2nd. Edition). Berlin : Springer.
Johnson, Diane (1999). Implementing a Hybrid Online Analytical Processing (HOLAP) solution. In Purba, Sanjiv (Ed.). Data Management Handbook. (3rd. edition, pp. 747-753). Boca Raton : Auerbach.
Lieberman, M., Ricciardi, T., Masarie, F., Spackman, K. (2003), The Use of SNOMED-CT Simplifies Querying of a Clinical Data Warehouse . AMIA Annu Symp Proc. 2003;
2003. 910. Retrieved November 16, 2006 from
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1480260
Meyer, Don, Cannon, Casey (1998). Building a Better Data Warehouse. Upper Saddle River : Prentice Hall.
Murphy, S., Gainer, V., Chueh, H.C. (2003), A Visual Interface Designed for Novice
Users to find Research Patient Cohorts in a Large Biomedical Database . AMIA
Annu Symp Proc. 2003;
2003. 489–493. Retrieved November 15, 2006 from
http://www.pubmedcentral.nih.gov/articlerender.fcgi?tool=pmcentrez&artid=1480150
Scalzo, Bert (2003). Oracle DBA Guide to Data Warehousing and Star Schemas.
Upper Saddle River : Prentice Hall PTR.
[r1] Do we have a reference made for this yet?
[r2] Marco, should we remove this? Does it just complicate matters? We didn’t do that because we are supporting 3 rule types (1to1 and architypes) as well… but that is somewhat off topic as it relates to warehouse architecture… thoughts? Rob
[r3] This is the first place we mention archetypes in the article. Should we introduce our 3 rules types somewhere? 1to1, aggregates and archetypes?
[r4] Mark, we should add a justification for your Mapped Term prefix here? Rob
[r5] Prakash has noted that instead of one Observable Fact View we may want to have a view for Observable Facts combined with the results of 1to1 mappings and a second View for aggregates and architypes. This is a pretty different View structure that would replace the Observable Fact View portion of the design with 2 Views. Rob is okay with that so long as we are able to make it work within the i2b2 Workbench. This issue will remain outstanding until we have had more prototyping and input on performance.