Page History
...
Highlight of Features
Top New Features
Contribution
Contributor
Description
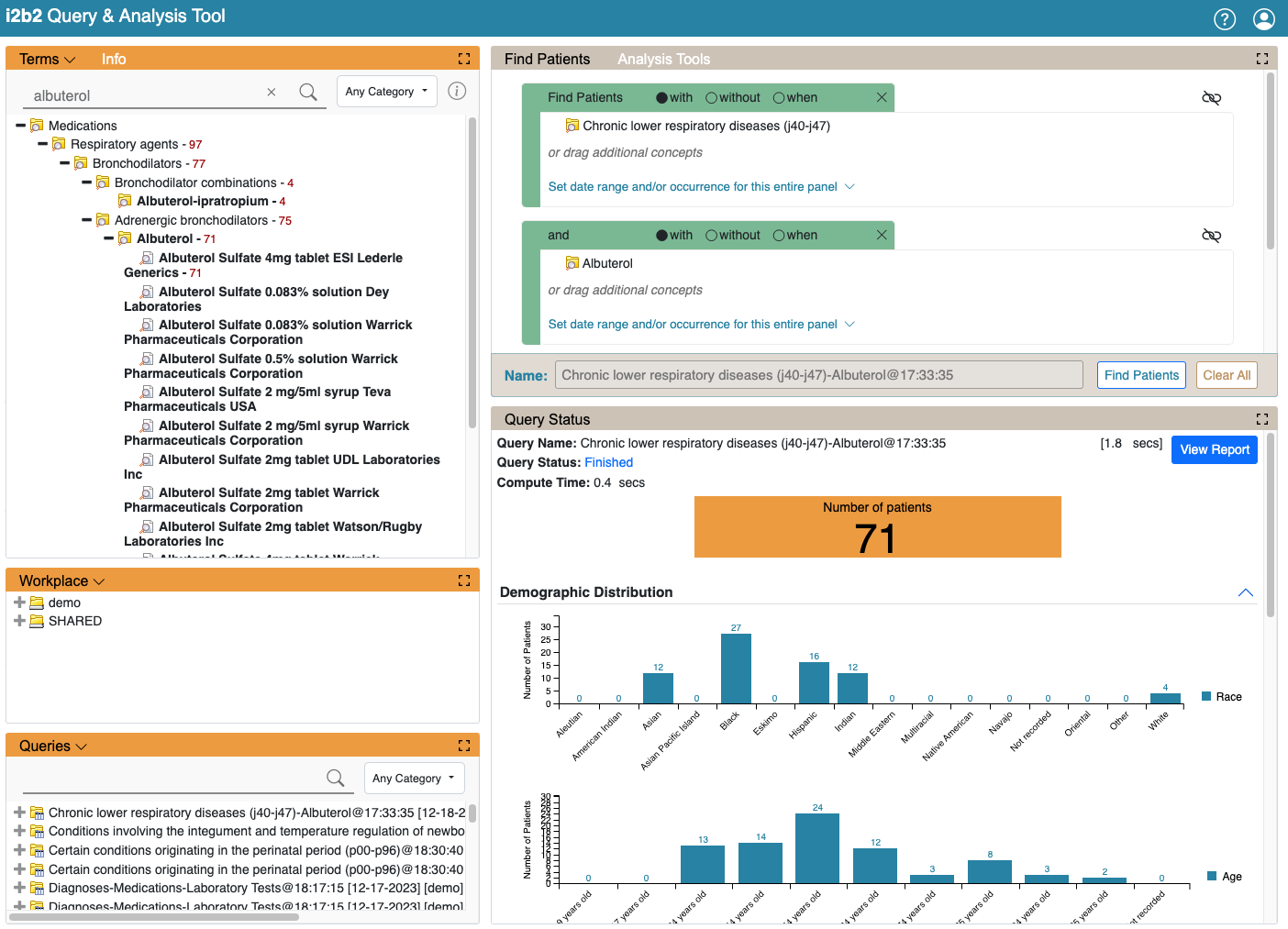
New Web client User Interface
Nick Benik (Harvard Medical School)
| Description | ||
|---|---|---|
New Web client User Interface | ACT i2b2 on OMOP for MSSQL and Oracle databases | |
Bug fixes |
Community-Contributed Features
The completely rewritten web client represents a major step forward for i2b2 that features an improved layout, greater visual customization, and a new plugin architecture. The new web client version eliminates usage of YUI |
and uses the latest versions of jQuery, Bootstrap 5 and Golden-Layout libraries, which will ensure maintainability far into the future. | ||
ACT i2b2 on OMOP for MSSQL and Oracle databases | The core i2b2 platform now supports the OMOP data model, which is queryable through the comprehensive ENACT Ontology. (MSSQL and Oracle are supported. Postgres support will be added in 1.8.1.) | |
Bug fixes |
Community-Contributed Features
Contribution | Contributor | Description |
| ACT i2b2 on OMOP | Michelle Morris (University of Pittsburgh) Mike Mendis, Jeff Klann, Reeta Metta (Mass General Brigham) | With i2b2 interface, using ACT-OMOP Ontology, queries can be run against OMOP data with OMOP table view structure. For information on installing and running queries using ACT-OMOP Data model |
Detailed Documentation on New Features
...
- Download and extract the new install zip package from "Download Binary Distribution" section of https://www.i2b2.org/software
- Edit the edu.harvard.i2b2.data\Release_1-7\NewInstall\Metadata\db.properties file to update the project properties to 'ACT-OMOP'; example: db.project=ACT-OMOP
- From the edu.harvard.i2b2.data\Release_1-7\NewInstall\Metadata folder, run the ant command: ant -f data_build.xml db_metadata_load_data
- This will execute the SQL scripts from the edu.harvard.i2b2.data\Release_1-7\NewInstall\Metadata\act\scripts\<db type> folder and will create and load ACT-OMOP Ontology metadata tables.
- You can now verify the new Ontology by logging into the web client.
Step 2: Load OMOP views and Concept dimension
- From the edu.harvard.i2b2.data\Release_1-7\NewInstall\Crcdata folder, run the ant command: ant -f data_build.xml db_demodata_load_data
- This will execute the BUILD_ACT_OMOP_Views _<dbtpye> script from the edu.harvard.i2b2.data\Release_1-7\NewInstall\crcdata\act-omop\scripts\<db type> folder and will create the OMOP views under the Views section in the database.
- In addition, the corresponding ACT-OMOP concept_dimension data is populated by executing the scripts form the crcdata.zip folders.
2. You can now run queries against the OMOP tables from the web client interface
| Note |
|---|
Delete your Patient_dimension and Visit_dimension tables before executing the crc scripts. The OMOP views will build these as Views. |
Step 3: Execute Totalnum counts
...
.svg.med.png)